Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T88451
|
|||||
| Target Name |
Mutated tyrosine-protein kinase Kit (mKIT)
|
|||||
| Synonyms |
v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog (mutated); p145 c-kit (mutated); Tyrosine-protein kinase Kit (mutated); SCFR; Proto-oncogene c-Kit (mutated); Piebald trait protein (mutated); PBT (mutated); Mast/stem cell growth factor receptor Kit (mutated); CD117 (mutated)
Click to Show/Hide
|
|||||
| Gene Name |
KIT
|
|||||
| Target Type |
Literature-reported target
|
[1] | ||||
| Function |
Tyrosine-protein kinase that acts as cell-surface receptor for the cytokine KITLG/SCF and plays an essential role in the regulation of cell survival and proliferation, hematopoiesis, stem cell maintenance, gametogenesis, mast cell development, migration and function, and in melanogenesis. In response to KITLG/SCF binding, KIT can activate several signaling pathways. Phosphorylates PIK3R1, PLCG1, SH2B2/APS and CBL. Activates the AKT1 signaling pathway by phosphorylation of PIK3R1, the regulatory subunit of phosphatidylinositol 3-kinase. Activated KIT also transmits signals via GRB2 and activation of RAS, RAF1 and the MAP kinases MAPK1/ERK2 and/or MAPK3/ERK1. Promotes activation of STAT family members STAT1, STAT3, STAT5A and STAT5B. Activation of PLCG1 leads to the production of the cellular signaling molecules diacylglycerol and inositol 1,4,5-trisphosphate. KIT signaling is modulated by protein phosphatases, and by rapid internalization and degradation of the receptor. Activated KIT promotes phosphorylation of the protein phosphatases PTPN6/SHP-1 and PTPRU, and of the transcription factors STAT1, STAT3, STAT5A and STAT5B. Promotes phosphorylation of PIK3R1, CBL, CRK (isoform Crk-II), LYN, MAPK1/ERK2 and/or MAPK3/ERK1, PLCG1, SRC and SHC1.
Click to Show/Hide
|
|||||
| BioChemical Class |
Kinase
|
|||||
| UniProt ID | ||||||
| EC Number |
EC 2.7.10.1
|
|||||
| Sequence |
MRGARGAWDFLCVLLLLLRVQTGSSQPSVSPGEPSPPSIHPGKSDLIVRVGDEIRLLCTD
PGFVKWTFEILDETNENKQNEWITEKAEATNTGKYTCTNKHGLSNSIYVFVRDPAKLFLV DRSLYGKEDNDTLVRCPLTDPEVTNYSLKGCQGKPLPKDLRFIPDPKAGIMIKSVKRAYH RLCLHCSVDQEGKSVLSEKFILKVRPAFKAVPVVSVSKASYLLREGEEFTVTCTIKDVSS SVYSTWKRENSQTKLQEKYNSWHHGDFNYERQATLTISSARVNDSGVFMCYANNTFGSAN VTTTLEVVDKGFINIFPMINTTVFVNDGENVDLIVEYEAFPKPEHQQWIYMNRTFTDKWE DYPKSENESNIRYVSELHLTRLKGTEGGTYTFLVSNSDVNAAIAFNVYVNTKPEILTYDR LVNGMLQCVAAGFPEPTIDWYFCPGTEQRCSASVLPVDVQTLNSSGPPFGKLVVQSSIDS SAFKHNGTVECKAYNDVGKTSAYFNFAFKGNNKEQIHPHTLFTPLLIGFVIVAGMMCIIV MILTYKYLQKPMYEVQWKVVEEINGNNYVYIDPTQLPYDHKWEFPRNRLSFGKTLGAGAF GKVVEATAYGLIKSDAAMTVAVKMLKPSAHLTEREALMSELKVLSYLGNHMNIVNLLGAC TIGGPTLVITEYCCYGDLLNFLRRKRDSFICSKQEDHAEAALYKNLLHSKESSCSDSTNE YMDMKPGVSYVVPTKADKRRSVRIGSYIERDVTPAIMEDDELALDLEDLLSFSYQVAKGM AFLASKNCIHRDLAARNILLTHGRITKICDFGLARDIKNDSNYVVKGNARLPVKWMAPES IFNCVYTFESDVWSYGIFLWELFSLGSSPYPGMPVDSKFYKMIKEGFRMLSPEHAPAEMY DIMKTCWDADPLKRPTFKQIVQLIEKQISESTNHIYSNLANCSPNRQKPVVDHSVRINSV GSTASSSQPLLVHDDV Click to Show/Hide
|
|||||
| Drug Binding Sites of Target | Top | |||||
|---|---|---|---|---|---|---|
| Ligand Name: Sunitinib | Ligand Info | |||||
| Structure Description | KIT kinase domain mutant D816H in complex with sunitinib | PDB:3G0F | ||||
| Method | X-ray diffraction | Resolution | 2.60 Å | Mutation | Yes | [2] |
| PDB Sequence |
NNYVYIDPTQ
575 LPYDHKWEFP585 RNRLSFGKTL595 GAGAFGKVVE605 ATAYGLIKSD615 AAMTVAVKML 625 KPSAHLTERE635 ALMSELKVLS645 YLGNHMNIVN655 LLGACTIGGP665 TLVITEYCCY 675 GDLLNFLRRK685 RDSFLDLEDL769 LSFSYQVAKG779 MAFLASKNCI789 HRDLAARNIL 799 LTHGRITKIC809 DFGLARHIKN819 DSNYVVKGNA829 RLPVKWMAPE839 SIFNCVYTFE 849 SDVWSYGIFL859 WELFSLGSSP869 YPGMPVDSKF879 YKMIKEGFRM889 LSPEHAPAEM 899 YDIMKTCWDA909 DPLKRPTFKQ919 IVQLIEKQIS929 E
|
|||||
|
|
||||||
| Ligand Name: adenosine diphosphate | Ligand Info | |||||
| Structure Description | Structure of a c-Kit Kinase Product Complex | PDB:1PKG | ||||
| Method | X-ray diffraction | Resolution | 2.90 Å | Mutation | Yes | [3] |
| PDB Sequence |
NVIDPTQLPY
578 DHKWEFPRNR588 LSFGKTLGAG598 AFGKVVEATA608 YGLIKSDAAM618 TVAVKMLKPS 628 AHLTEREALM638 SELKVLSYLG648 NHMNIVNLLG658 ACTIGGPTLV668 ITEYCCYGDL 678 LNFLRRKRDS688 FICSKALDLE767 DLLSFSYQVA777 KGMAFLASKN787 CIHRDLAARN 797 ILLTHGRITK807 ICDFGLARDI817 KNDSNYVVKG827 NARLPVKWMA837 PESIFNCVYT 847 FESDVWSYGI857 FLWELFSLGS867 SPYPGMPVDS877 KFYKMIKEGF887 RMLSPEHAPA 897 EMYDIMKTCW907 DADPLKRPTF917 KQIVQLIEKQ927
|
|||||
|
|
LEU595
3.748
GLY596
3.561
ALA597
3.944
GLY598
2.674
ALA599
3.428
PHE600
3.290
GLY601
2.858
LYS602
4.946
VAL603
3.474
ALA621
3.381
LYS623
2.709
GLU640
4.588
|
|||||
| Click to View More Binding Site Information of This Target with Different Ligands | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Human Pathway Affiliation
|
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
| MAPK signaling pathway | hsa04010 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Ras signaling pathway | hsa04014 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Rap1 signaling pathway | hsa04015 | Affiliated Target |
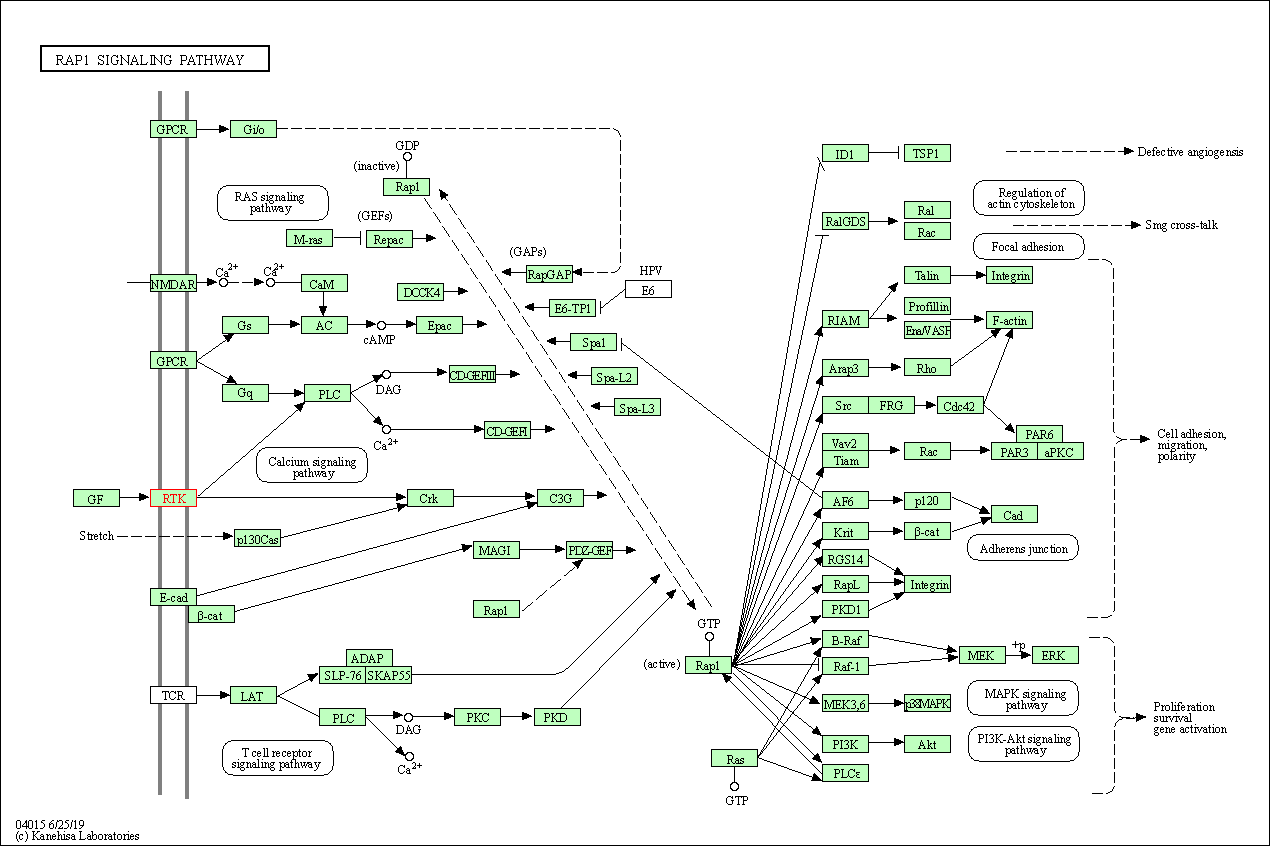
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Phospholipase D signaling pathway | hsa04072 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| PI3K-Akt signaling pathway | hsa04151 | Affiliated Target |
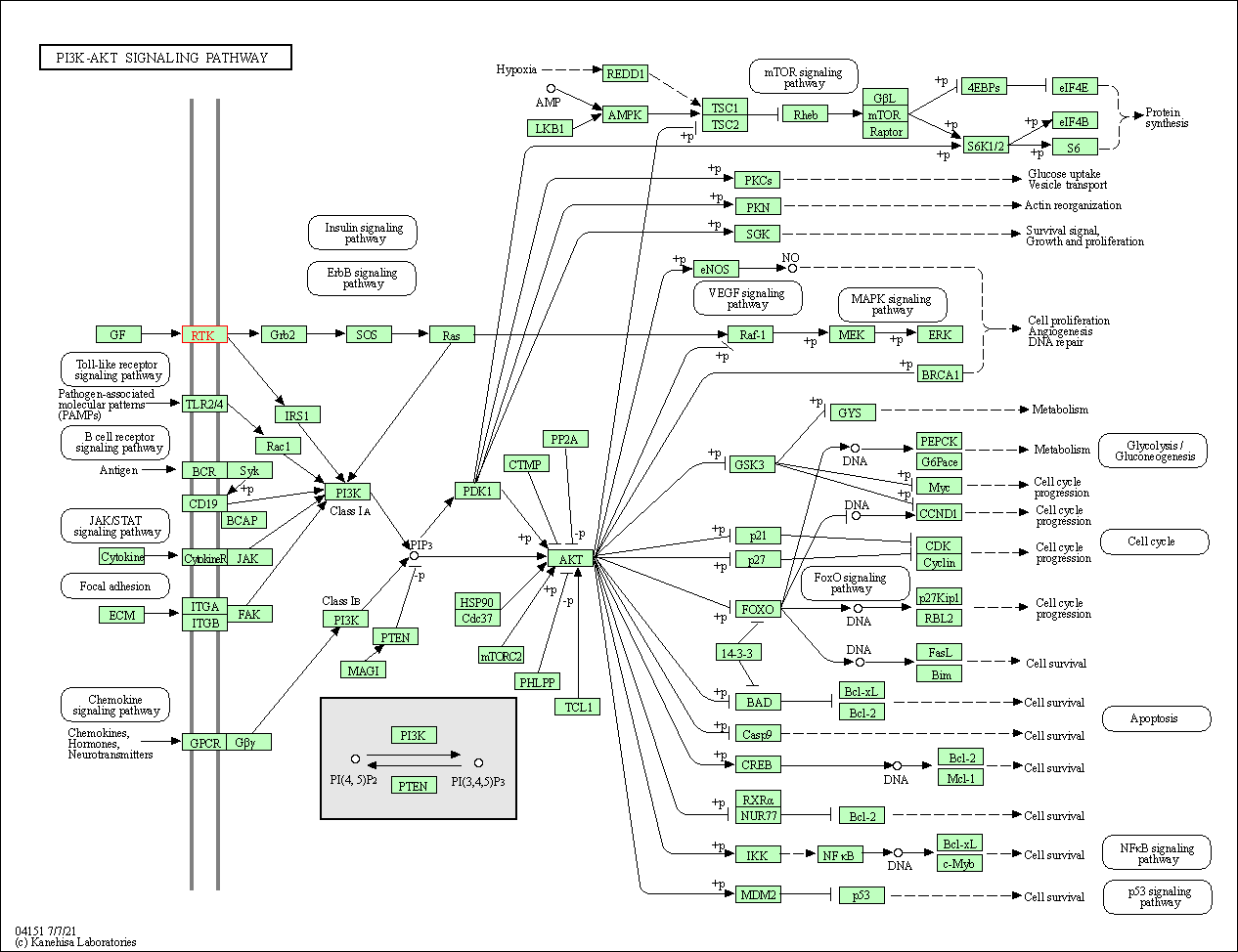
|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Hematopoietic cell lineage | hsa04640 | Affiliated Target |
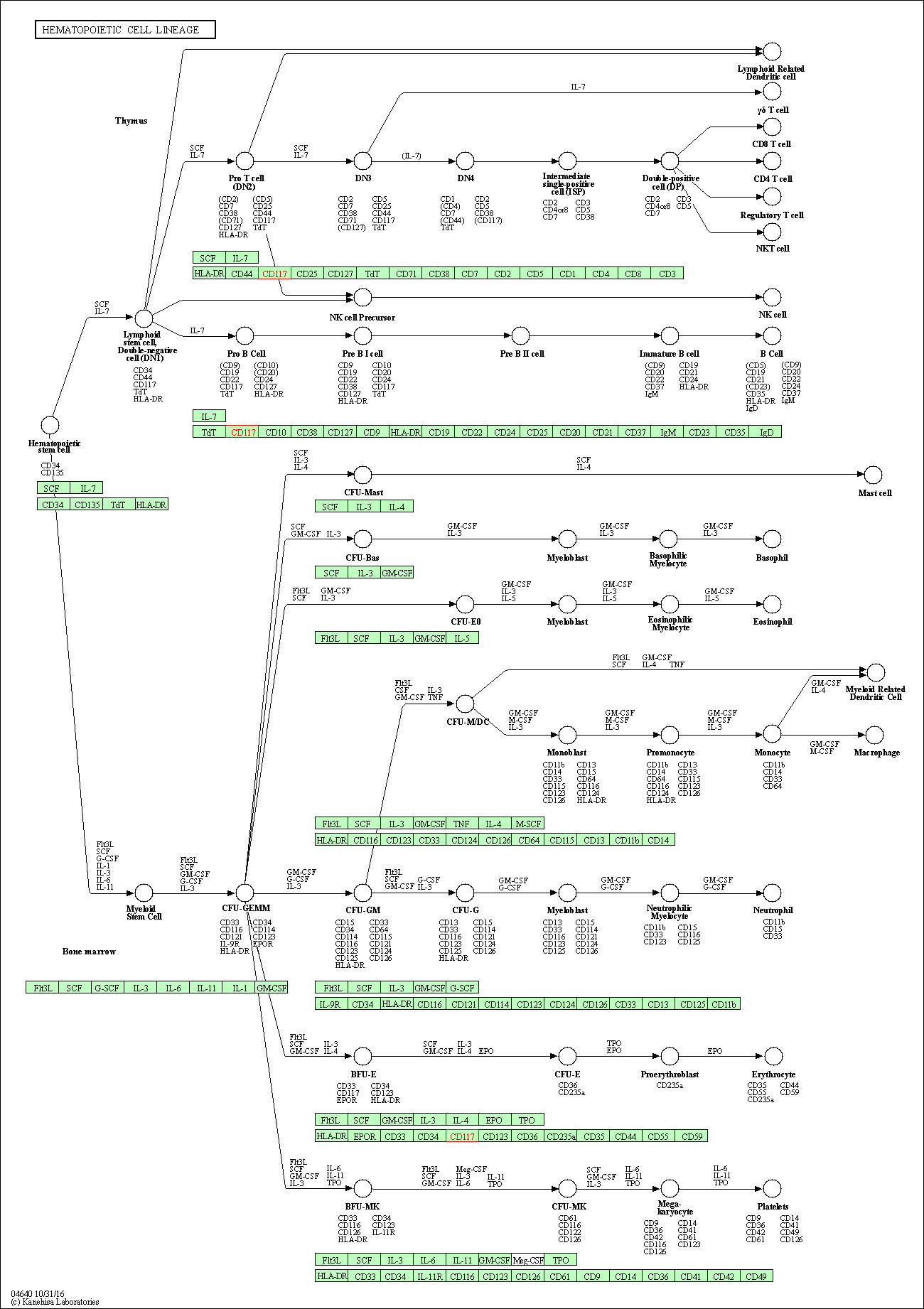
|
| Class: Organismal Systems => Immune system | Pathway Hierarchy | ||
| Melanogenesis | hsa04916 | Affiliated Target |
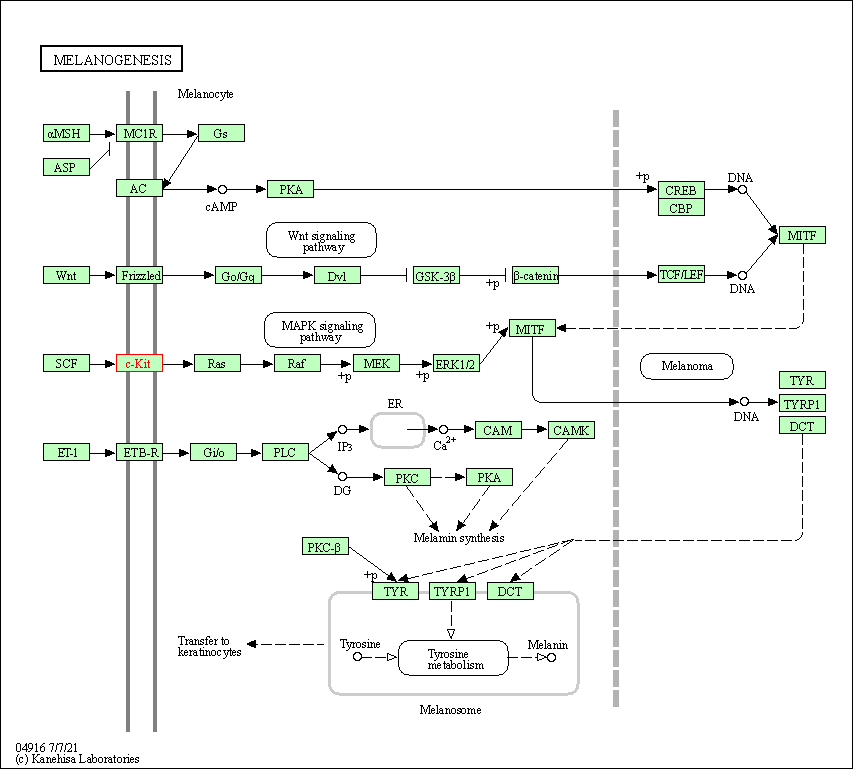
|
| Class: Organismal Systems => Endocrine system | Pathway Hierarchy | ||
| Click to Show/Hide the Information of Affiliated Human Pathways | |||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Clinical pipeline report, company report or official report of the Pharmaceutical Research and Manufacturers of America (PhRMA) | |||||
| REF 2 | KIT kinase mutants show unique mechanisms of drug resistance to imatinib and sunitinib in gastrointestinal stromal tumor patients. Proc Natl Acad Sci U S A. 2009 Feb 3;106(5):1542-7. | |||||
| REF 3 | Structure of a c-kit product complex reveals the basis for kinase transactivation. J Biol Chem. 2003 Aug 22;278(34):31461-4. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

