| Drug General Information |
|---|
| Drug ID |
D07HVY |
| Drug Name |
Efavirenz |
Drug Info
|
| Synonyms |
EFV; EFZ; Eravirenz; Stocrin; Sustiva; DMP 266; L 743726; Atripla (TN); DMP-266; L-741211; L-743725; L-743726; Stocrin (TN); Strocin (TM); Sustiva (TM); Sustiva (TN); Efavirenz (JAN/INN); L-743,726; Zoxazin-2-one; Efavirenz, (S)-isomer; Met-SDF-1.beta. & Efavirenz; Met-Stromal Cell-derived Factor-1.beta. (Human) & Efavirenz; (-)-Efavirenz; (4S)-6-Chloro-4-(cyclopropylethynyl)-1,4-dihydro-4-(trifluoromethyl)-2H-3,1-benzoxazin-2-one; (4S)-6-Chloro-4-cyclopropylethynyl-4-trifluoromethyl-1,4-dihydro-benzo[d][1,3]oxazin-2-one; (4S)-6-chloro-4-(2-cyclopropylethynyl)-4-(trifluoromethyl)-1H-3,1-benzoxazin-2-one; (4S)-6-chloro-4-(cyclopropylethynyl)-4-(trifluoromethyl)-1,4-dihydro-2H-3,1-benzoxazin-2-one; (S)-6-Chloro-4-(2-cyclopropylethynyl)-1,4-dihydro-4-(trifluoromethyl)-2H-3,1-ben; (S)-6-Chloro-4-(cyclopropylethynyl)-1,4-dihydro-4-(trifluoromethyl)-2H-3,1-benzoxazin-2-one; (S)-6-Chloro-4-cyclopropylethynyl-4-trifluoromethyl-1,4-dihydro-benzo[d][1,3]oxazin-2-one; 2H-3,1-Benzoxazin-2-one, 6-chloro-4-(cyclopropylethynyl)-1,4-dihydro-4-(trifluoromethyl)-, (4S)-(9; 6-chloro-4-(2-cyclopropyl-1-ethynyl)-4-trifluoromethyl-(4S)-1,4-dihydro-2H-benzo[d][1,3]oxazin-2-one; EFV |
| Drug Type |
Small molecular drug |
| Therapeutic Class |
Anti-HIV Agents |
| Structure |
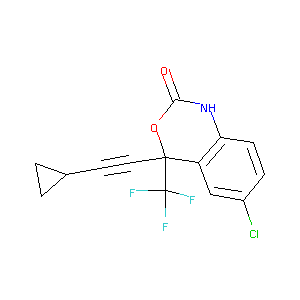
|
| Drug Resistance Mutations |
|---|
| Target Name |
HIV Non-Nucleoside reverse transcriptase |
|
| Uniprot ID |
POL_HV1B1(600-1159) |
| Species |
Human immunodeficiency virus type 1 (HIV-1) |
| Reference Sequence |
PISPIETVPVKLKPGMDGPKVKQWPLTEEKIKALVEICTEMEKEGKISKIGPENPYNTPV
FAIKKKDSTKWRKLVDFRELNKRTQDFWEVQLGIPHPAGLKKKKSVTVLDVGDAYFSVPL
DEDFRKYTAFTIPSINNETPGIRYQYNVLPQGWKGSPAIFQSSMTKILEPFKKQNPDIVI
YQYMDDLYVGSDLEIGQHRTKIEELRQHLLRWGLTTPDKKHQKEPPFLWMGYELHPDKWT
VQPIVLPEKDSWTVNDIQKLVGKLNWASQIYPGIKVRQLCKLLRGTKALTEVIPLTEEAE
LELAENREILKEPVHGVYYDPSKDLIAEIQKQGQGQWTYQIYQEPFKNLKTGKYARMRGA
HTNDVKQLTEAVQKITTESIVIWGKTPKFKLPIQKETWETWWTEYWQATWIPEWEFVNTP
PLVKLWYQLEKEPIVGAETFYVDGAANRETKLGKAGYVTNKGRQKVVPLTNTTNQKTELQ
AIYLALQDSGLEVNIVTDSQYALGIIQAQPDKSESELVNQIIEQLIKKEKVYLAWVPAHK
GIGGNEQVDKLVSAGIRKIL [Human immunodeficiency virus type 1 (H
IV-1)] |
| Targeted Disease |
HIV infection |
| Drug Resistance Mutations |
| Mutation info |
Missense: Y188L |
[555484]
|
| Level of Resistance |
Reduce EFV susceptibility 5 fold |
|
| Mutation info |
Missense: K103N |
[555479]
|
| Level of Resistance |
Reduce EFV susceptibility about 20 fold |
|
| Mutation info |
Missense: V106A |
[556164]
|
| Level of Resistance |
Reduce EFV susceptibility 5 fold |
|
| Mutation info |
Missense: K101P |
[555757]
|
| Level of Resistance |
Reduce EFV susceptibility >50 fold |
|
| Mutation info |
Missense: L100I |
[555479]
|
| Level of Resistance |
High-level resistance |
|
| Mutation info |
Missense: Y181C |
[555757]
|
| Level of Resistance |
Reduce EFV susceptibility about 2 fold |
|
| Mutation info |
Missense: V108I |
[555887]
|
| Level of Resistance |
Reduce EFV susceptibility about 2 fold |
|
| Mutation info |
Missense: A98G |
[555544],
[555618],
[555925]
|
| Level of Resistance |
Reduce EFV susceptibility about 2-3 fold |
|
| Mutation info |
Missense: N348I |
[555649],
[555651],
[555751]
|
| Level of Resistance |
Reduce EFV susceptibility 2 fold |
|
| Mutation info |
Missense: K101E |
[555479],
[555757],
[555773]
|
| Level of Resistance |
Reduce EFV susceptibility 1-5 fold |
|
| Mutation info |
Missense: G190E |
[555572],
[555737],
[555925]
|
| Level of Resistance |
Confer high-level resistance to EFV |
|
| Mutation info |
Missense: V179E |
[555510],
[555701],
[555925]
|
| Level of Resistance |
Confer low-level resistance to EFV |
|
| Mutation info |
Missense: G190A |
[555509],
[555544],
[555618]
|
| Level of Resistance |
Reduce EFV susceptibility 5-10 fold |
|
| Mutation info |
Missense: G190S |
[555509],
[555544],
[555618]
|
| Level of Resistance |
Reduce EFV susceptibility >50 fold |
|
| Mutation info |
Missense: K103S |
[555563],
[555618],
[555962]
|
| Level of Resistance |
Cause high-level resistance to EFV |
|
| Mutation info |
Missense: K238T |
[555510],
[555581],
[555925]
|
| Level of Resistance |
Reduce EFV susceptibility about 5 fold |
|
| Mutation info |
Missense: G190Q |
[555509],
[555962]
|
| Level of Resistance |
Confer high-level resistance to EFV |
|
| Mutation info |
Missense: Y188H |
[555618],
[555962]
|
| Level of Resistance |
Reduce EFV susceptibility 5 fold |
|
| Mutation info |
Missense: V179D |
[555701],
[555962]
|
| Level of Resistance |
Reduce EFV susceptibility 2-5 fold |
|
| Mutation info |
Missense: K238N |
[555726],
[555925]
|
| Level of Resistance |
Reduce susceptibility to EFV |
|
| Mutation info |
Missense: L100V |
[555914],
[555925]
|
| Level of Resistance |
Reduce EFV susceptibility 10 fold |
|
| Mutation info |
Missense: V106M |
[555925],
[555962]
|
| Level of Resistance |
Reduce EFV susceptibility >30 fold |
|
| Mutation info |
Missense: F227L |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: G190C |
[556228],
[556246]
|
| Level of Resistance |
High-level resistance |
|
| Mutation info |
Missense: G190T |
[556228],
[556246]
|
| Level of Resistance |
High-level resistance |
|
| Mutation info |
Missense: G190V |
[556228],
[556246]
|
| Level of Resistance |
High-level resistance |
|
| Mutation info |
Missense: K103R + V179D |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: K103T |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: M230L |
[556228],
[556246]
|
| Level of Resistance |
Intermediate resistance |
|
| Mutation info |
Missense: P225H |
[556228],
[556246]
|
| Level of Resistance |
Intermediate resistance |
|
| Mutation info |
Missense: V106A + F227L |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: Y181F |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: Y181G |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: Y181I |
[556228],
[556246]
|
| Level of Resistance |
Intermediate resistance |
|
| Mutation info |
Missense: Y181S |
[556228],
[556246]
|
| Level of Resistance |
Low-level resistance |
|
| Mutation info |
Missense: Y181V |
[556228],
[556246]
|
| Level of Resistance |
Intermediate resistance |
|
| Mutation info |
Missense: Y188F |
[556228],
[556246]
|
| Level of Resistance |
High-level resistance |
|
| Mutation info |
Missense: K103H |
[555563]
|
| Level of Resistance |
Reduce EFV susceptibility 20 fold |
|
| Mutation info |
Missense: Y188C |
[555962]
|
| Level of Resistance |
Reduce EFV susceptibility 20 fold |
|
| Mutation info |
Missense: E138G |
[556038]
|
| Level of Resistance |
Reduce EFV susceptibility 2 fold |
|
| Mutation info |
Missense: F227C |
[556038]
|
| Level of Resistance |
Reduce EFV susceptibility 4-5 fold |
|
| Mutation info |
Missense: M230I |
[556038]
|
| Level of Resistance |
Reduce EFV susceptibility 11 fold |
|
| References |
|---|
| Ref 550326 | Approved antiretroviral drugs. Antiretroviral Drugs. Company report of AVERT. 2009. |
|---|
| Ref 555479 | Human immunodeficiency virus type 1 mutations selected in patients failing efavirenz combination therapy. Antimicrob Agents Chemother. 2000 Sep;44(9):2475-84. |
|---|
| Ref 555484 | Genotypic correlates of phenotypic resistance to efavirenz in virus isolates from patients failing nonnucleoside reverse transcriptase inhibitor therapy. J Virol. 2001 Jun;75(11):4999-5008. |
|---|
| Ref 555509 | Amino acid substitutions at position 190 of human immunodeficiency virus type 1 reverse transcriptase increase susceptibility to delavirdine and impair virus replication. J Virol. 2003 Jan;77(2):1512-23. |
|---|
| Ref 555510 | Human immunodeficiency virus reverse transcriptase and protease sequence database. Nucleic Acids Res. 2003 Jan 1;31(1):298-303. |
|---|
| Ref 555544 | Distribution of human immunodeficiency virus type 1 protease and reverse transcriptase mutation patterns in 4,183 persons undergoing genotypic resistance testing. Antimicrob Agents Chemother. 2004 Aug;48(8):3122-6. |
|---|
| Ref 555563 | Rare mutations at codon 103 of HIV-1 reverse transcriptase can confer resistance to non-nucleoside reverse transcriptase inhibitors. AIDS. 2005 Mar 24;19(6):549-54. |
|---|
| Ref 555572 | TMC125 displays a high genetic barrier to the development of resistance: evidence from in vitro selection experiments. J Virol. 2005 Oct;79(20):12773-82. |
|---|
| Ref 555581 | The K101P and K103R/V179D mutations in human immunodeficiency virus type 1 reverse transcriptase confer resistance to nonnucleoside reverse transcriptase inhibitors. Antimicrob Agents Chemother. 2006 Jan;50(1):351-4. |
|---|
| Ref 555618 | A novel nonnucleoside analogue that inhibits human immunodeficiency virus type 1 isolates resistant to current nonnucleoside reverse transcriptase inhibitors. Antimicrob Agents Chemother. 2007 Feb;51(2):429-37. Epub 2006 Nov 20. |
|---|
| Ref 555649 | N348I in the connection domain of HIV-1 reverse transcriptase confers zidovudine and nevirapine resistance. PLoS Med. 2007 Dec;4(12):e335. |
|---|
| Ref 555651 | Amino acid mutation N348I in the connection subdomain of human immunodeficiency virus type 1 reverse transcriptase confers multiclass resistance to nucleoside and nonnucleoside reverse transcriptase inhibitors. J Virol. 2008 Apr;82(7):3261-70. doi: 10.1128/JVI.01154-07. Epub 2008 Jan 23. |
|---|
| Ref 555701 | Compilation and prevalence of mutations associated with resistance to non-nucleoside reverse transcriptase inhibitors. Antivir Ther. 2009;14(1):103-9. |
|---|
| Ref 555726 | Nonpolymorphic human immunodeficiency virus type 1 protease and reverse transcriptase treatment-selected mutations. Antimicrob Agents Chemother. 2009 Nov;53(11):4869-78. doi: 10.1128/AAC.00592-09. Epub 2009 Aug 31. |
|---|
| Ref 555737 | TMC278, a next-generation nonnucleoside reverse transcriptase inhibitor (NNRTI), active against wild-type and NNRTI-resistant HIV-1. Antimicrob Agents Chemother. 2010 Feb;54(2):718-27. doi: 10.1128/AAC.00986-09. Epub 2009 Nov 23. |
|---|
| Ref 555751 | Combinations of mutations in the connection domain of human immunodeficiency virus type 1 reverse transcriptase: assessing the impact on nucleoside and nonnucleoside reverse transcriptase inhibitor resistance. Antimicrob Agents Chemother. 2010 May;54(5):1973-80. doi: 10.1128/AAC.00870-09. Epub 2010 Mar 1. |
|---|
| Ref 555757 | Constrained patterns of covariation and clustering of HIV-1 non-nucleoside reverse transcriptase inhibitor resistance mutations. J Antimicrob Chemother. 2010 Jul;65(7):1477-85. doi: 10.1093/jac/dkq140. Epub 2010 May 12. |
|---|
| Ref 555773 | Characterization of genotypic and phenotypic changes in HIV-1-infected patients with virologic failure on an etravirine-containing regimen in the DUET-1 and DUET-2 clinical studies. AIDS Res Hum Retroviruses. 2010 Nov;26(11):1197-205. doi: 10.1089/aid.2009.0302. Epub 2010 Sep 20. |
|---|
| Ref 556228 | The HIVdb system for HIV-1 genotypic resistance interpretation. Intervirology. 2012;55(2):98-101. doi: 10.1159/000331998. Epub 2012 Jan 24. |
|---|
| Ref 555887 | Identification of drug resistant mutations in HIV-1 CRF07_BC variants selected by nevirapine in vitro. PLoS One. 2012;7(9):e44333. doi: 10.1371/journal.pone.0044333. Epub 2012 Sep 12. |
|---|
| Ref 555914 | Quantitative prediction of integrase inhibitor resistance from genotype through consensus linear regression modeling. Virol J. 2013 Jan 3;10:8. doi: 10.1186/1743-422X-10-8. |
|---|
| Ref 555925 | Significantly improved HIV inhibitor efficacy prediction employing proteochemometric models generated from antivirogram data. PLoS Comput Biol. 2013;9(2):e1002899. doi: 10.1371/journal.pcbi.1002899. Epub 2013 Feb 21. |
|---|
| Ref 555962 | Non-nucleoside reverse transcriptase inhibitor (NNRTI) cross-resistance: implications for preclinical evaluation of novel NNRTIs and clinical genotypic resistance testing. J Antimicrob Chemother. 2014 Jan;69(1):12-20. doi: 10.1093/jac/dkt316. Epub 2013 Aug 9. |
|---|
| Ref 556038 | Impact of drug resistance-associated amino acid changes in HIV-1 subtype C on susceptibility to newer nonnucleoside reverse transcriptase inhibitors. Antimicrob Agents Chemother. 2015 Feb;59(2):960-71. doi: 10.1128/AAC.04215-14. Epub 2014 Nov 24. |
|---|
| Ref 556246 | Development and significance of nucleoside drug resistance in infection caused by the human immunodeficiency virus type 1. Clin Invest Med. 1994 Jun;17(3):226-43. |
|---|
| Ref 556164 | Convergent combination therapy can select viable multidrug-resistant HIV-1 in vitro. Nature. 1993 Sep 30;365(6445):451-3. |
|---|


