Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T04364
(Former ID: TTDI00246)
|
|||||
| Target Name |
Cullin-3 (CUL-3)
|
|||||
| Synonyms |
KIAA0617; Cullin3
Click to Show/Hide
|
|||||
| Gene Name |
CUL3
|
|||||
| Target Type |
Literature-reported target
|
[1] | ||||
| Function |
BCR complexes and ARIH1 collaborate in tandem to mediate ubiquitination of target proteins. As a scaffold protein may contribute to catalysis through positioning of the substrate and the ubiquitin-conjugating enzyme. The E3 ubiquitin-protein ligase activity of the complex is dependent on the neddylation of the cullin subunit and is inhibited by the association of the deneddylated cullin subunit with TIP120A/CAND1. The functional specificity of the BCR complex depends on the BTB domain-containing protein as the substrate recognition component. BCR(KLHL42) is involved in ubiquitination of KATNA1. BCR(SPOP) is involved in ubiquitination of BMI1/PCGF4, BRMS1, H2AFY and DAXX, GLI2 and GLI3. Can also form a cullin-RING-based BCR (BTB-CUL3-RBX1) E3 ubiquitin-protein ligase complex containing homodimeric SPOPL or the heterodimer formed by SPOP and SPOPL; these complexes have lower ubiquitin ligase activity. BCR(KLHL9-KLHL13) controls the dynamic behavior of AURKB on mitotic chromosomes and thereby coordinates faithful mitotic progression and completion of cytokinesis. BCR(KLHL12) is involved in ER-Golgi transport by regulating the size of COPII coats, thereby playing a key role in collagen export, which is required for embryonic stem (ES) cells division: BCR(KLHL12) acts by mediating monoubiquitination of SEC31 (SEC31A or SEC31B). BCR(KLHL3) acts as a regulator of ion transport in the distal nephron; by mediating ubiquitination of WNK4. The BCR(KLHL20) E3 ubiquitin ligase complex is involved in interferon response and anterograde Golgi to endosome transport: it mediates both ubiquitination leading to degradation and 'Lys-33'-linked ubiquitination. The BCR(KLHL21) E3 ubiquitin ligase complex regulates localization of the chromosomal passenger complex (CPC) from chromosomes to the spindle midzone in anaphase and mediates the ubiquitination of AURKB. The BCR(KLHL22) ubiquitin ligase complex mediates monoubiquitination of PLK1, leading to PLK1 dissociation from phosphoreceptor proteins and subsequent removal from kinetochores, allowing silencing of the spindle assembly checkpoint (SAC) and chromosome segregation. The BCR(KLHL22) ubiquitin ligase complex is also responsible for the amino acid-stimulated 'Lys-48' polyubiquitination and proteasomal degradation of DEPDC5. Through the degradation of DEPDC5, releases the GATOR1 complex-mediated inhibition of the TORC1 pathway. The BCR(KLHL25) ubiquitin ligase complex is involved in translational homeostasis by mediating ubiquitination and subsequent degradation of hypophosphorylated EIF4EBP1 (4E-BP1). The BCR(KBTBD8) complex acts by mediating monoubiquitination of NOLC1 and TCOF1, leading to remodel the translational program of differentiating cells in favor of neural crest specification. Involved in ubiquitination of cyclin E and of cyclin D1 (in vitro) thus involved in regulation of G1/S transition. Involved in the ubiquitination of KEAP1, ENC1 and KLHL41. In concert with ATF2 and RBX1, promotes degradation of KAT5 thereby attenuating its ability to acetylate and activate ATM. The BCR(KCTD17) E3 ubiquitin ligase complex mediates ubiquitination and degradation of TCHP, a down-regulator of cilium assembly, thereby inducing ciliogenesis. The BCR(KLHL24) E3 ubiquitin ligase complex mediates ubiquitination of KRT14, controls KRT14 levels during keratinocytes differentiation, and is essential for skin integrity. Core component of multiple cullin-RING-based BCR (BTB-CUL3-RBX1) E3 ubiquitin-protein ligase complexes which mediate the ubiquitination and subsequent proteasomal degradation of target proteins.
Click to Show/Hide
|
|||||
| UniProt ID | ||||||
| Sequence |
MSNLSKGTGSRKDTKMRIRAFPMTMDEKYVNSIWDLLKNAIQEIQRKNNSGLSFEELYRN
AYTMVLHKHGEKLYTGLREVVTEHLINKVREDVLNSLNNNFLQTLNQAWNDHQTAMVMIR DILMYMDRVYVQQNNVENVYNLGLIIFRDQVVRYGCIRDHLRQTLLDMIARERKGEVVDR GAIRNACQMLMILGLEGRSVYEEDFEAPFLEMSAEFFQMESQKFLAENSASVYIKKVEAR INEEIERVMHCLDKSTEEPIVKVVERELISKHMKTIVEMENSGLVHMLKNGKTEDLGCMY KLFSRVPNGLKTMCECMSSYLREQGKALVSEEGEGKNPVDYIQGLLDLKSRFDRFLLESF NNDRLFKQTIAGDFEYFLNLNSRSPEYLSLFIDDKLKKGVKGLTEQEVETILDKAMVLFR FMQEKDVFERYYKQHLARRLLTNKSVSDDSEKNMISKLKTECGCQFTSKLEGMFRDMSIS NTTMDEFRQHLQATGVSLGGVDLTVRVLTTGYWPTQSATPKCNIPPAPRHAFEIFRRFYL AKHSGRQLTLQHHMGSADLNATFYGPVKKEDGSEVGVGGAQVTGSNTRKHILQVSTFQMT ILMLFNNREKYTFEEIQQETDIPERELVRALQSLACGKPTQRVLTKEPKSKEIENGHIFT VNDQFTSKLHRVKIQTVAAKQGESDPERKETRQKVDDDRKHEIEAAIVRIMKSRKKMQHN VLVAEVTQQLKARFLPSPVVIKKRIEGLIEREYLARTPEDRKVYTYVA Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | AlphaFold | ||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
|
There is no similarity protein (E value < 0.005) for this target
|
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
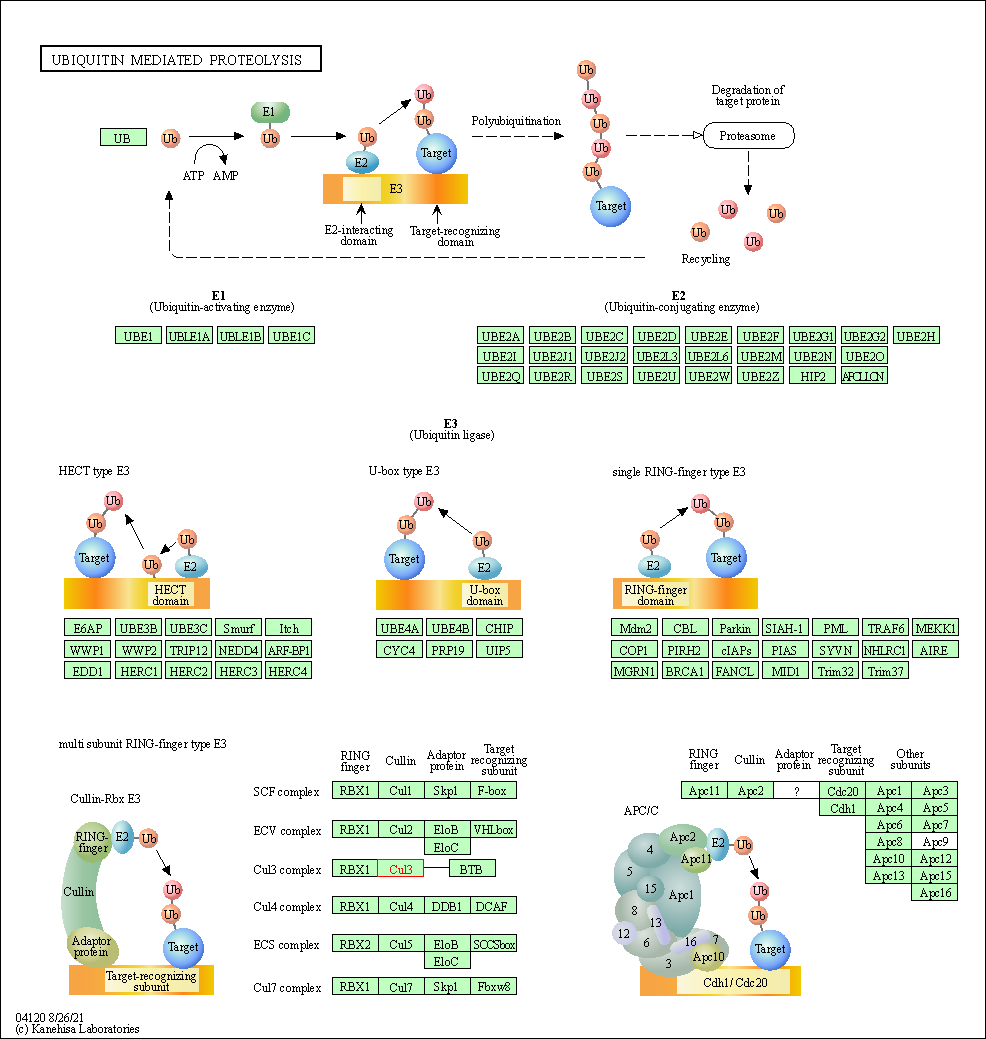
| Ubiquitin mediated proteolysis | hsa04120 | Affiliated Target |

|
| Class: Genetic Information Processing => Folding, sorting and degradation | Pathway Hierarchy | ||
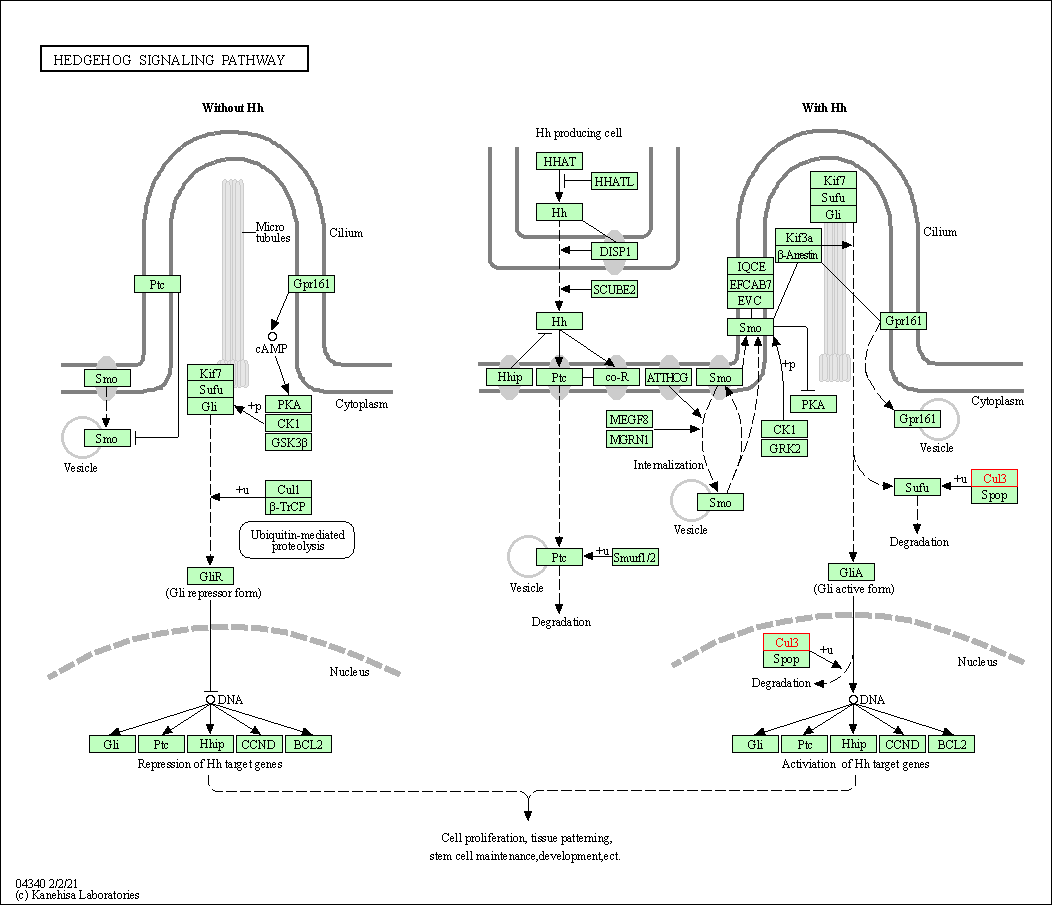
| Hedgehog signaling pathway | hsa04340 | Affiliated Target |

|
| Class: Environmental Information Processing => Signal transduction | Pathway Hierarchy | ||
| Degree | 51 | Degree centrality | 5.48E-03 | Betweenness centrality | 6.36E-03 |
|---|---|---|---|---|---|
| Closeness centrality | 2.25E-01 | Radiality | 1.40E+01 | Clustering coefficient | 2.75E-02 |
| Neighborhood connectivity | 9.10E+00 | Topological coefficient | 3.20E-02 | Eccentricity | 12 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Target Regulators | Top | |||||
|---|---|---|---|---|---|---|
| Target-interacting Proteins | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Functional analysis of Cullin 3 E3 ligases in tumorigenesis. Biochim Biophys Acta Rev Cancer. 2018 Jan;1869(1):11-28. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

