Target Information
| Target General Information | Top | |||||
|---|---|---|---|---|---|---|
| Target ID |
T29678
(Former ID: TTDI01968)
|
|||||
| Target Name |
Voltage-gated sodium channel alpha Nav1.2 (SCN2A)
|
|||||
| Synonyms |
Voltage-gated sodium channelsubunit alpha Nav1.2; Sodium channel protein type II subunit alpha; Sodium channel protein brainII subunit alpha; SCN2A; HBSC II
Click to Show/Hide
|
|||||
| Gene Name |
SCN2A
|
|||||
| Target Type |
Literature-reported target
|
[1] | ||||
| Function |
Mediates the voltage-dependent sodium ion permeability of excitable membranes. Assuming opened or closed conformations in response to the voltage difference across the membrane, the protein forms a sodium-selective channel through which Na(+) ions may pass in accordance with their electrochemical gradient.
Click to Show/Hide
|
|||||
| BioChemical Class |
Voltage-gated ion channel
|
|||||
| UniProt ID | ||||||
| Sequence |
MAQSVLVPPGPDSFRFFTRESLAAIEQRIAEEKAKRPKQERKDEDDENGPKPNSDLEAGK
SLPFIYGDIPPEMVSVPLEDLDPYYINKKTFIVLNKGKAISRFSATPALYILTPFNPIRK LAIKILVHSLFNMLIMCTILTNCVFMTMSNPPDWTKNVEYTFTGIYTFESLIKILARGFC LEDFTFLRDPWNWLDFTVITFAYVTEFVDLGNVSALRTFRVLRALKTISVIPGLKTIVGA LIQSVKKLSDVMILTVFCLSVFALIGLQLFMGNLRNKCLQWPPDNSSFEINITSFFNNSL DGNGTTFNRTVSIFNWDEYIEDKSHFYFLEGQNDALLCGNSSDAGQCPEGYICVKAGRNP NYGYTSFDTFSWAFLSLFRLMTQDFWENLYQLTLRAAGKTYMIFFVLVIFLGSFYLINLI LAVVAMAYEEQNQATLEEAEQKEAEFQQMLEQLKKQQEEAQAAAAAASAESRDFSGAGGI GVFSESSSVASKLSSKSEKELKNRRKKKKQKEQSGEEEKNDRVRKSESEDSIRRKGFRFS LEGSRLTYEKRFSSPHQSLLSIRGSLFSPRRNSRASLFSFRGRAKDIGSENDFADDEHST FEDNDSRRDSLFVPHRHGERRHSNVSQASRASRVLPILPMNGKMHSAVDCNGVVSLVGGP STLTSAGQLLPEGTTTETEIRKRRSSSYHVSMDLLEDPTSRQRAMSIASILTNTMEELEE SRQKCPPCWYKFANMCLIWDCCKPWLKVKHLVNLVVMDPFVDLAITICIVLNTLFMAMEH YPMTEQFSSVLSVGNLVFTGIFTAEMFLKIIAMDPYYYFQEGWNIFDGFIVSLSLMELGL ANVEGLSVLRSFRLLRVFKLAKSWPTLNMLIKIIGNSVGALGNLTLVLAIIVFIFAVVGM QLFGKSYKECVCKISNDCELPRWHMHDFFHSFLIVFRVLCGEWIETMWDCMEVAGQTMCL TVFMMVMVIGNLVVLNLFLALLLSSFSSDNLAATDDDNEMNNLQIAVGRMQKGIDFVKRK IREFIQKAFVRKQKALDEIKPLEDLNNKKDSCISNHTTIEIGKDLNYLKDGNGTTSGIGS SVEKYVVDESDYMSFINNPSLTVTVPIAVGESDFENLNTEEFSSESDMEESKEKLNATSS SEGSTVDIGAPAEGEQPEVEPEESLEPEACFTEDCVRKFKCCQISIEEGKGKLWWNLRKT CYKIVEHNWFETFIVFMILLSSGALAFEDIYIEQRKTIKTMLEYADKVFTYIFILEMLLK WVAYGFQVYFTNAWCWLDFLIVDVSLVSLTANALGYSELGAIKSLRTLRALRPLRALSRF EGMRVVVNALLGAIPSIMNVLLVCLIFWLIFSIMGVNLFAGKFYHCINYTTGEMFDVSVV NNYSECKALIESNQTARWKNVKVNFDNVGLGYLSLLQVATFKGWMDIMYAAVDSRNVELQ PKYEDNLYMYLYFVIFIIFGSFFTLNLFIGVIIDNFNQQKKKFGGQDIFMTEEQKKYYNA MKKLGSKKPQKPIPRPANKFQGMVFDFVTKQVFDISIMILICLNMVTMMVETDDQSQEMT NILYWINLVFIVLFTGECVLKLISLRYYYFTIGWNIFDFVVVILSIVGMFLAELIEKYFV SPTLFRVIRLARIGRILRLIKGAKGIRTLLFALMMSLPALFNIGLLLFLVMFIYAIFGMS NFAYVKREVGIDDMFNFETFGNSMICLFQITTSAGWDGLLAPILNSGPPDCDPDKDHPGS SVKGDCGNPSVGIFFFVSYIIISFLVVVNMYIAVILENFSVATEESAEPLSEDDFEMFYE VWEKFDPDATQFIEFAKLSDFADALDPPLLIAKPNKVQLIAMDLPMVSGDRIHCLDILFA FTKRVLGESGEMDALRIQMEERFMASNPSKVSYEPITTTLKRKQEEVSAIIIQRAYRRYL LKQKVKKVSSIYKKDKGKECDGTPIKEDTLIDKLNENSTPEKTDMTPSTTSPPSYDSVTK PEKEKFEKDKSEKEDKGKDIRESKK Click to Show/Hide
|
|||||
| 3D Structure | Click to Show 3D Structure of This Target | PDB | ||||
| Cell-based Target Expression Variations | Top | |||||
|---|---|---|---|---|---|---|
| Cell-based Target Expression Variations | ||||||
| Drug Binding Sites of Target | Top | |||||
|---|---|---|---|---|---|---|
| Ligand Name: (3beta,14beta,17beta,25R)-3-[4-methoxy-3-(methoxymethyl)butoxy]spirost-5-en | Ligand Info | |||||
| Structure Description | Human Nav1.2-beta2-KIIIA ternary complex | PDB:6J8E | ||||
| Method | Electron microscopy | Resolution | 3.00 Å | Mutation | No | [3] |
| PDB Sequence |
PIRKLAIKIL
126 VHSLFNMLIM136 CTILTNCVFM146 TMSNPPDWTK156 NVEYTFTGIY166 TFESLIKILA 176 RGFCLEDFTF186 LRDPWNWLDF196 TVITFAYVTE206 FVDLGNVSAL216 RTFRVLRALK 226 TISVIPGLKT236 IVGALIQSVK246 KLSDVMILTV256 FCLSVFALIG266 LQLFMGNLRN 276 KCLQWPPDFN315 WDEYIEDKSH325 FYFLEGQNDA335 LLCGNSSDAG345 QCPEGYICVK 355 AGRNPNYGYT365 SFDTFSWAFL375 SLFRLMTQDF385 WENLYQLTLR395 AAGKTYMIFF 405 VLVIFLGSFY415 LINLILAVVA425 MAYEEQNQAT435 LEEAEQDCCK743 PWLKVKHLVN 753 LVVMDPFVDL763 AITICIVLNT773 LFMAMEHYPM783 TEQFSSVLSV793 GNLVFTGIFT 803 AEMFLKIIAM813 DPYYYFQEGW823 NIFDGFIVSL833 SLMELGLANV843 EGLSVLRSFR 853 LLRVFKLAKS863 WPTLNMLIKI873 IGNSVGALGN883 LTLVLAIIVF893 IFAVVGMQLF 903 GKSYKECVCK913 ISNDCELPRW923 HMHDFFHSFL933 IVFRVLCGEW943 IETMWDCMEV 953 AGQTMCLTVF963 MMVMVIGNLV973 VLNLFLALLL983 SSFSGKLWWN1196 LRKTCYKIVE 1206 HNWFETFIVF1216 MILLSSGALA1226 FEDIYIEQRK1236 TIKTMLEYAD1246 KVFTYIFILE 1256 MLLKWVAYGF1266 QVYFTNAWCW1276 LDFLIVDVSL1286 VSLTANALGY1296 SELGAIKSLR 1306 TLRALRPLRA1316 LSRFEGMRVV1326 VNALLGAIPS1336 IMNVLLVCLI1346 FWLIFSIMGV 1356 NLFAGKFYHC1366 INYTTGEMFD1376 VSVVNNYSEC1386 KALIESNQTA1396 RWKNVKVNFD 1406 NVGLGYLSLL1416 QVATFKGWMD1426 IMYAAVDSRN1436 VELQPKYEDN1446 LYMYLYFVIF 1456 IIFGSFFTLN1466 LFIGVIIDNF1476 NQQKKKFGGQ1486 DIFMTEEQKK1496 YYNAMKKLGS 1506 KKPQKPIPRP1516 ANKFQGMVFD1526 FVTKQVFDIS1536 IMILICLNMV1546 TMMVETDDQS 1556 QEMTNILYWI1566 NLVFIVLFTG1576 ECVLKLISLR1586 YYYFTIGWNI1596 FDFVVVILSI 1606 VGMFLAELIE1616 KYFVSPTLFR1626 VIRLARIGRI1636 LRLIKGAKGI1646 RTLLFALMMS 1656 LPALFNIGLL1666 LFLVMFIYAI1676 FGMSNFAYVK1686 REVGIDDMFN1696 FETFGNSMIC 1706 LFQITTSAGW1716 DGLLAPILNS1726 GPPDCDPDKD1736 HPGSSVKGDC1746 GNPSVGIFFF 1756 VSYIIISFLV1766 VVNMYIAVIL1776 ENFSVATEE
|
|||||
|
|
LEU421
3.333
ALA425
3.644
GLU429
4.794
PHE978
3.661
LEU979
3.555
LEU982
3.619
LEU983
3.727
PHE986
4.075
LEU1465
4.174
ILE1469
4.059
|
|||||
| Click to View More Binding Site Information of This Target with Different Ligands | ||||||
| Different Human System Profiles of Target | Top |
|---|---|
|
Human Similarity Proteins
of target is determined by comparing the sequence similarity of all human proteins with the target based on BLAST. The similarity proteins for a target are defined as the proteins with E-value < 0.005 and outside the protein families of the target.
A target that has fewer human similarity proteins outside its family is commonly regarded to possess a greater capacity to avoid undesired interactions and thus increase the possibility of finding successful drugs
(Brief Bioinform, 21: 649-662, 2020).
Human Tissue Distribution
of target is determined from a proteomics study that quantified more than 12,000 genes across 32 normal human tissues. Tissue Specificity (TS) score was used to define the enrichment of target across tissues.
The distribution of targets among different tissues or organs need to be taken into consideration when assessing the target druggability, as it is generally accepted that the wider the target distribution, the greater the concern over potential adverse effects
(Nat Rev Drug Discov, 20: 64-81, 2021).
Human Pathway Affiliation
of target is determined by the life-essential pathways provided on KEGG database. The target-affiliated pathways were defined based on the following two criteria (a) the pathways of the studied target should be life-essential for both healthy individuals and patients, and (b) the studied target should occupy an upstream position in the pathways and therefore had the ability to regulate biological function.
Targets involved in a fewer pathways have greater likelihood to be successfully developed, while those associated with more human pathways increase the chance of undesirable interferences with other human processes
(Pharmacol Rev, 58: 259-279, 2006).
Biological Network Descriptors
of target is determined based on a human protein-protein interactions (PPI) network consisting of 9,309 proteins and 52,713 PPIs, which were with a high confidence score of ≥ 0.95 collected from STRING database.
The network properties of targets based on protein-protein interactions (PPIs) have been widely adopted for the assessment of target’s druggability. Proteins with high node degree tend to have a high impact on network function through multiple interactions, while proteins with high betweenness centrality are regarded to be central for communication in interaction networks and regulate the flow of signaling information
(Front Pharmacol, 9, 1245, 2018;
Curr Opin Struct Biol. 44:134-142, 2017).
Human Similarity Proteins
Human Tissue Distribution
Human Pathway Affiliation
Biological Network Descriptors
|
|
| Protein Name | Pfam ID | Percentage of Identity (%) | E value |
|---|---|---|---|
| Sodium/hydrogen exchanger 11 (SLC9C2) | 27.273 (36/132) | 3.37E-04 |
|
Note:
If a protein has TS (tissue specficity) scores at least in one tissue >= 2.5, this protein is called tissue-enriched (including tissue-enriched-but-not-specific and tissue-specific). In the plots, the vertical lines are at thresholds 2.5 and 4.
|
| KEGG Pathway | Pathway ID | Affiliated Target | Pathway Map |
|---|---|---|---|
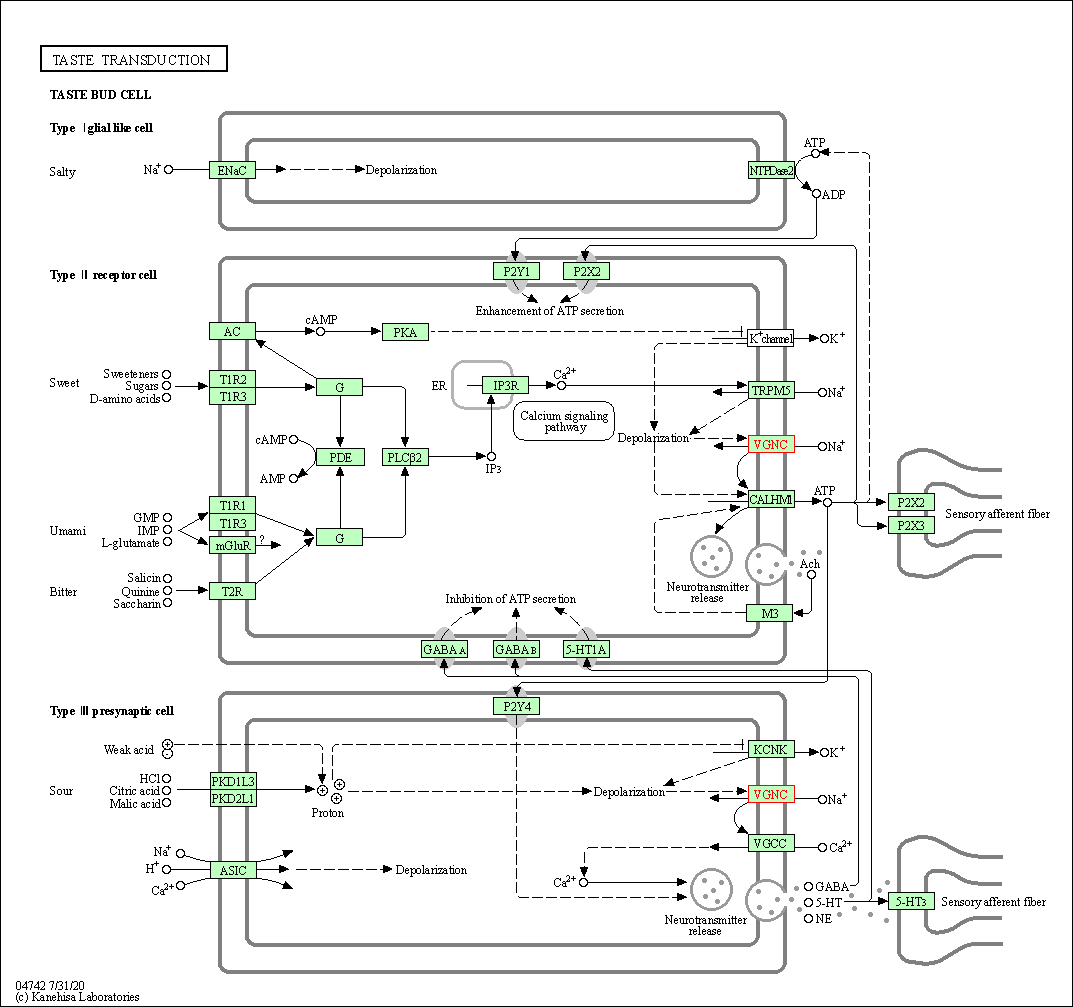
| Taste transduction | hsa04742 | Affiliated Target |

|
| Class: Organismal Systems => Sensory system | Pathway Hierarchy | ||
| Degree | 3 | Degree centrality | 3.22E-04 | Betweenness centrality | 2.03E-04 |
|---|---|---|---|---|---|
| Closeness centrality | 1.52E-01 | Radiality | 1.20E+01 | Clustering coefficient | 0.00E+00 |
| Neighborhood connectivity | 4.67E+00 | Topological coefficient | 3.67E-01 | Eccentricity | 14 |
| Download | Click to Download the Full PPI Network of This Target | ||||
| Chemical Structure based Activity Landscape of Target | Top |
|---|---|
| Drug Property Profile of Target | Top | |
|---|---|---|
| (1) Molecular Weight (mw) based Drug Clustering | (2) Octanol/Water Partition Coefficient (xlogp) based Drug Clustering | |
|
|
||
| (3) Hydrogen Bond Donor Count (hbonddonor) based Drug Clustering | (4) Hydrogen Bond Acceptor Count (hbondacc) based Drug Clustering | |
|
|
||
| (5) Rotatable Bond Count (rotbonds) based Drug Clustering | (6) Topological Polar Surface Area (polararea) based Drug Clustering | |
|
|
||
| "RO5" indicates the cutoff set by lipinski's rule of five; "D123AB" colored in GREEN denotes the no violation of any cutoff in lipinski's rule of five; "D123AB" colored in PURPLE refers to the violation of only one cutoff in lipinski's rule of five; "D123AB" colored in BLACK represents the violation of more than one cutoffs in lipinski's rule of five | ||
| Target Poor or Non Binders | Top | |||||
|---|---|---|---|---|---|---|
| Target Poor or Non Binders | ||||||
| References | Top | |||||
|---|---|---|---|---|---|---|
| REF 1 | Binding of batrachotoxinin A 20-alpha-benzoate to a receptor site associated with sodium channels in synaptic nerve ending particles. J Biol Chem. 1981 Sep 10;256(17):8922-7. | |||||
| REF 2 | Interaction of batrachotoxin with the local anesthetic receptor site in transmembrane segment IVS6 of the voltage-gated sodium channel. Proc Natl Acad Sci U S A. 1998 Nov 10;95(23):13947-52. | |||||
| REF 3 | Molecular basis for pore blockade of human Na(+) channel Na(v)1.2 by the u-conotoxin KIIIA. Science. 2019 Mar 22;363(6433):1309-1313. | |||||
If You Find Any Error in Data or Bug in Web Service, Please Kindly Report It to Dr. Zhou and Dr. Zhang.

